I recently saw some nice presentations of protein structures prepared by MOLMOL. Since MOLMOL is not actively maintained, I wonder how to make similar colors on the secondary structural elements in PyMOL.
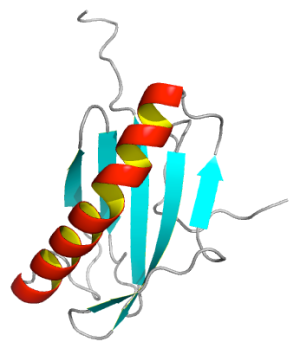
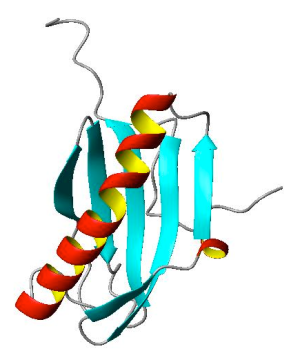
In this short blog, an example is shown. The top and bottom figures are made by PyMOL and MOLMOL, respectively.
How I did that? Basically, just make highlight color yellow, turn on discrete cartoon color, adjust the width and length of strands.
The PyMOL script is shown here, too:
cmd.show_as(“cartoon” ,”all”)
bg_color white
set cartoon_highlight_color, yellow
set cartoon_rect_width, 0.03
set cartoon_rect_length, 1.7
set cartoon_discrete_colors, on
util.cbss(“all”,”red”,”cyan”,”gray70″,_self=cmd)
Leave a comment